
Predicting the native or near-native binding pose of a small molecule within a protein binding pocket is an extremely important task in structure-based drug design, especially in the hit-to-lead and lead optimization phases. This enables us to provide relevant recommendations within the data-driven research life cycle to develop and/or procure an effective RDM system following the FAIR guiding principles. To understand the current state of data-driven materials science and learn about the challenges faced with RDM, we gather user stories of researchers from small- and large-scale projects. Hence, there are efforts to standardize, curate, preserve, and disseminate these data in a way that is findable, accessible, interoperable, and reusable (FAIR). The materials science community is no exception to these challenges as it heralds its new paradigm of data-driven science, which uses artificial intelligence to accelerate materials discovery but requires massive datasets to perform effectively. This may be attributed to the 5V challenge in big data: volume, variety, velocity, veracity, and value. 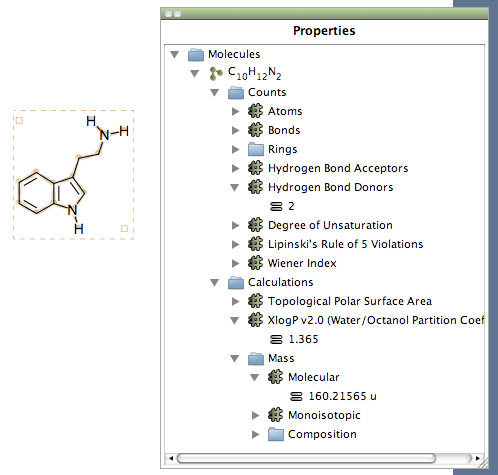
The need for good research data management (RDM) practices is becoming more recognized as a critical part of research. The comprehensive annotated information could give users a clear picture of their de novo library and could guide the further selection of candidates for chemical synthesis and biological confirmation. DenovoProfiling could provide structural identification, chemical space exploration, drug mapping, and target & pathway information. Currently, DenovoProfiling contains six modules: (1) identification & visualization module for chemical structure visualization and identify the reported structures, (2) chemical space module for chemical space exploration using similarity maps, principal components analysis (PCA), drug-like properties distribution, and scaffold-based clustering, (3) ADMET prediction module for predicting the ADMET properties of the de novo molecules, (4) molecular alignment module for three dimensional molecular shape analysis, (5) drugs mapping module for identifying structural similar drugs, and (6) target & pathway module for identifying the reported targets and corresponding functional pathways. 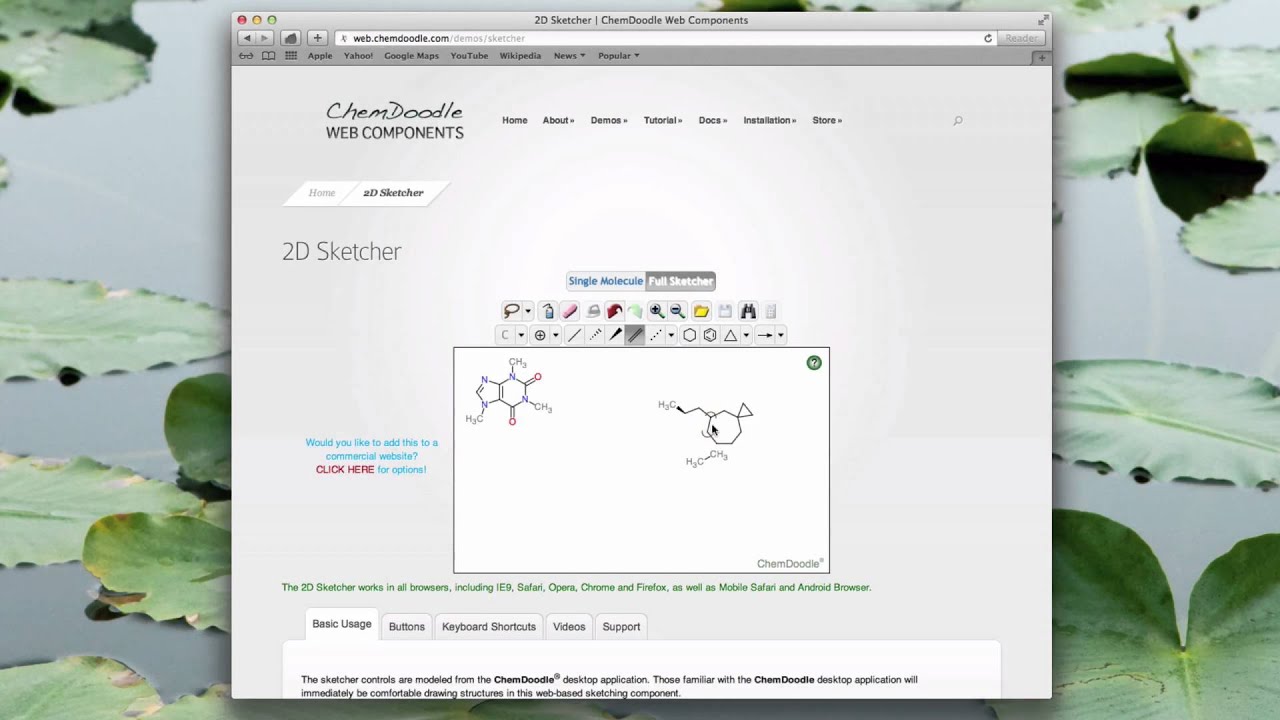
In this paper, we present DenovoProfiling, a webserver dedicated to de novo library visualization and functional profiling. An increasing number of publicly available chemogenomic databases sets good foundations and creates good opportunities for comprehensive profiling of the de novo library. The flourish of the de novo molecular generation methods and applications has created a great demand for the visualization and functional profiling for the de novo generated molecules. Various deep learning-based architectures for molecular generation have been proposed for de novo drug design. The plugin ( ) is easily adaptable to any academic user both embedding and similarity measures can be configured. This workflow is implemented as a Moodle plugin, using the Chemdoodle engine for drawing structures and communicating with a REST server to compute the similarity score using molecular descriptors. To overcome this limitation, we have developed a grading workflow based on the pairwise similarity score of two considered chemical structures. Specifically, a strict comparison of candidate and expected chemical structures is not sufficient when some tolerance is deemed acceptable. This is particularly true in the case of chemical structures, where most questions simply cannot be evaluated on a true/false basis. Therefore, such platforms are unevenly adapted to different disciplines. Typically, existing online platforms use a binary grading system, which often fails to give a nuanced evaluation of the answers given by the students. 
We report a novel approach for grading chemical structure drawings for remote teaching, integrated into the Moodle platform.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
